Welcome to the NoMoRe server!
What is Normal Mode Refinement?
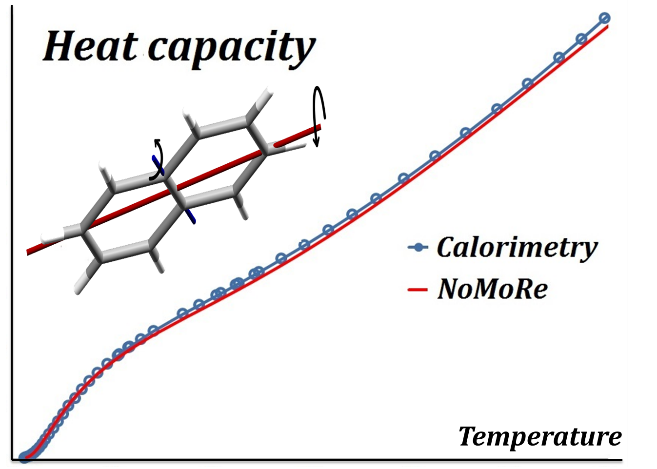
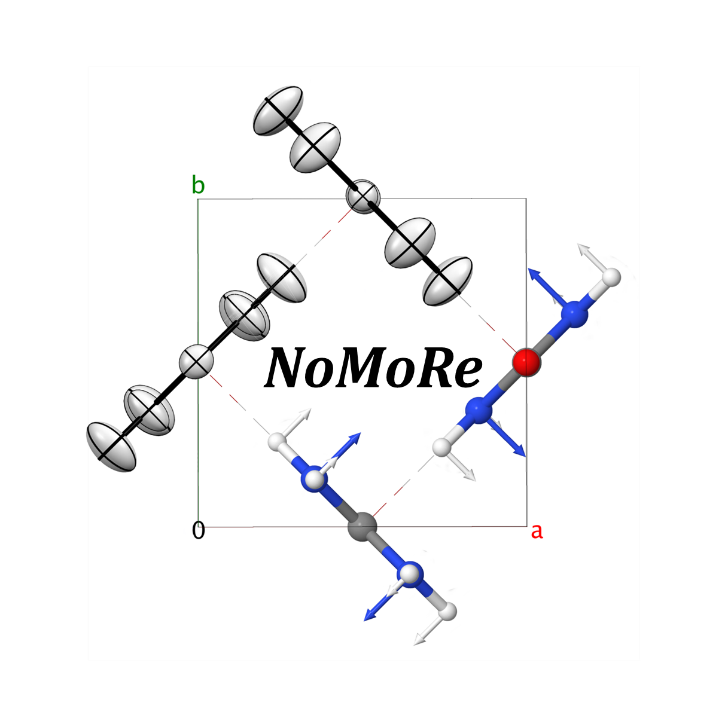
Normal Mode Refinement (NoMoRe) is a modelling approach which enables the refinement of frequencies of normal modes evaluated using ab-initio periodic computations against single crystal diffraction data. In this approach instead of refining anisotropic displacement parameters (ADPs) as in routine X-ray refinement, we are refining only few frequencies related to low-frequency modes, which corresponds to the external vibrations of molecule. Schematic workflow of our method is presented in Scheme 1.
The frequencies, which are obtained after the refinement, enable calculation of thermodynamic properties.
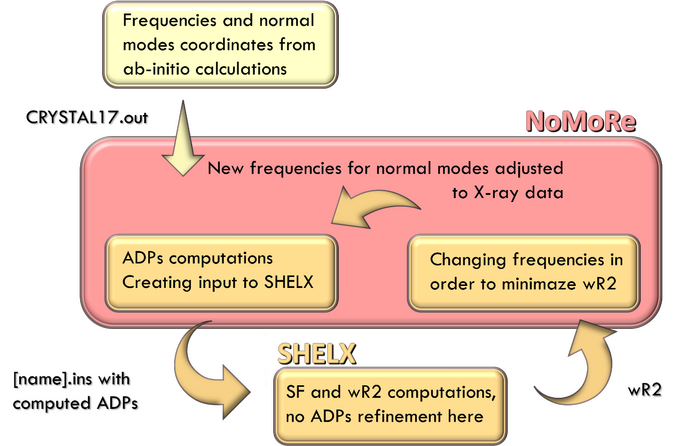
Scheme 1. Schematic workflow with NoMoRe.
What are the advantages of using NoMoRe?
- With fewer parameters we can obtain results comparable to standard ADP refinements.
- Refined frequencies enable evaluation of thermodynamic properties.
- H-atom ADPs agree well with neutron values.
Combination of NoMoRe with Aspherical Atom Models (AAM_NoMoRe)
Previously, when using NoMoRe, electron density was described by IAM. In following stage, we modified this method and held the refinement against aspherical atomic form factors (see Scheme 2). To do that we replaced shelxl with a DiSCaMB library based program which calculates structure factors using atomic form factors file in tsc format as an input. The tsc file is a table of scattering or form factors for each atom type. It can be generated with NoSpherA2, the implementation of HAR in OLEX2 (NoSpherA2: Non-Spherical Atoms in Olex2).

Scheme 2. Schematic workflow with AAM_NoMoRe.
Aspherical Atom Models
Aspherical atom models (AAM), such as HAR and TAAM, overcome limitations such as Hirshfeld atom refinement (HAR) and the Transferable Aspherical Atom Model (TAAM), overcome the limitations of simplified spherical approximations. This results in significantly enhanced accuracy and precision for hydrogen atom positions and their ADPs in single-crystal X-ray results compared to IAM. The concept of TAAM is based on the principle that multipolar parameters derived from the Hansen–Coppens model for atoms in one chemical environment can be applied to another similar environment, as the differences between them are effectively negligible. TAAM is based on databanks of different types of pseudoatoms which are constrained to predefined values characteristic for the corresponding atom type. HAR uses tailor-made aspherical atomic structure factors directly from quantum chemical calculations.
For more details see:
NoMoRe
- Hoser A. A., Madsen A. Ø., Acta Cryst A 2016, 72, 206–214.
- Hoser A. A., Madsen A. Ø., Acta Cryst A 2017, 73, 102–114.
- Kofoed P. M. et al., IUCrJ 2019.
AAM_NoMoRe
- Butkiewicz H. et al., IUCrJ 2025, 12, 123–136.
Aspherical Atom Models
- Jayatilaka D. & Dittrich B., Acta Cryst A 2008, 64, 383–393.
- Capelli S. C. et al., IUCrJ 2014, 1, 361–379.
- Pichon-Pesme V. et al., J. Phys. Chem. 1995, 99, 6242–6250.
- Jha K. K. et al., Acta Cryst B 2020, 76, 621–625.
How to run NoMoRe?
Because calculations can be time-consuming we decided to create an accounts for users. After calculations, the results will be available from the user account in the dashboard section.
Plese create an account and sign in with Login/Sign up section on the top right corner of screen.
After login in the menu three new options will appear:
- Dashboard – download results files
- Refinements – submit NoMoRe or HAR_NoMoRe
- Thermodynamics – compute Heat Capacity, Entropy, Enthalpy, Gibbs Free Energy
Required files for NoMoRe
- Gamma-point normal-mode results (CRYSTAL09/14/17/23) – input via cif2crystal.
- SHELX structure-factor file (
.hkl). - SHELX input (
.ins).
Additional files for AAM_NoMoRe
- Structure-factor file (
.fcf). - Form-factor table (
.tsc). - CIF structure file (
.cif).
All files must use Unix format LF.
Examples: NoMoRe examples | AAM_NoMoRe examples
Manual: download PDF
Important
This server is in development. Your comments and bug reports will be greatly appreciated and we will try to respond quickly with an updated version.
We recommend that you critically inspect the resulting ADPs using a visualization program, e.g. Ortep, Platon, Mercury or Peanut.
Contact: nomore_help@chem.uw.edu.pl
Contact
Anna A. Hoser
University of Warsaw, Chemistry Department
a.hoser@uw.edu.pl
Profile
Anders Ø. Madsen
University of Copenhagen, Dept. of Pharmacy
a.madsen@sund.ku.dk
Profile
Funding
- Foundation for Polish Science
- Villum Foundation